There is a family tree rooted at 0 consisting of n nodes numbered 0 to n - 1. You are given a 0-indexed integer array parents, where parents[i] is the parent for node i. Since node 0 is the root, parents[0] == -1.
There are 105 genetic values, each represented by an integer in the inclusive range [1, 105]. You are given a 0-indexed integer array nums, where nums[i] is a distinct genetic value for node i.
Return an array ans of length n where ans[i] is the smallest genetic value that is missing from the subtree rooted at node i.
The subtree rooted at a node x contains node x and all of its descendant nodes.
Example 1:
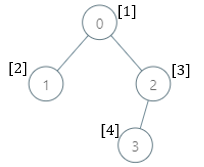
Input: parents = [-1,0,0,2], nums = [1,2,3,4] Output: [5,1,1,1] Explanation: The answer for each subtree is calculated as follows: - 0: The subtree contains nodes [0,1,2,3] with values [1,2,3,4]. 5 is the smallest missing value. - 1: The subtree contains only node 1 with value 2. 1 is the smallest missing value. - 2: The subtree contains nodes [2,3] with values [3,4]. 1 is the smallest missing value. - 3: The subtree contains only node 3 with value 4. 1 is the smallest missing value.
Example 2:
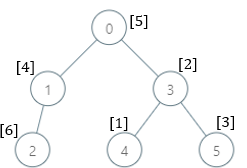
Input: parents = [-1,0,1,0,3,3], nums = [5,4,6,2,1,3] Output: [7,1,1,4,2,1] Explanation: The answer for each subtree is calculated as follows: - 0: The subtree contains nodes [0,1,2,3,4,5] with values [5,4,6,2,1,3]. 7 is the smallest missing value. - 1: The subtree contains nodes [1,2] with values [4,6]. 1 is the smallest missing value. - 2: The subtree contains only node 2 with value 6. 1 is the smallest missing value. - 3: The subtree contains nodes [3,4,5] with values [2,1,3]. 4 is the smallest missing value. - 4: The subtree contains only node 4 with value 1. 2 is the smallest missing value. - 5: The subtree contains only node 5 with value 3. 1 is the smallest missing value.
Example 3:
Input: parents = [-1,2,3,0,2,4,1], nums = [2,3,4,5,6,7,8] Output: [1,1,1,1,1,1,1] Explanation: The value 1 is missing from all the subtrees.
Constraints:
n == parents.length == nums.length2 <= n <= 1050 <= parents[i] <= n - 1 for i != 0parents[0] == -1parents represents a valid tree.1 <= nums[i] <= 105nums[i] is distinct.Problem Overview: You are given a rooted tree where each node has a unique genetic value. For every node, compute the smallest positive genetic value that does not appear in its subtree. The challenge is efficiently tracking which values exist inside each subtree without recomputing sets from scratch.
Approach 1: DFS with a Set to Track Genetic Values (O(n^2) worst-case time, O(n) space)
This method performs a depth-first search and builds a set of genetic values for each subtree. When DFS visits a node, it recursively collects the sets from its children and merges them into the current node’s set. After merging, iterate from value 1 upward until you find the smallest number not present in the set. That value becomes the answer for the current node. The key operation is merging child sets and performing hash lookups to detect missing values. This approach is easy to implement and clearly demonstrates subtree aggregation, but repeated set merges can degrade to O(n^2) time in skewed trees.
Approach 2: Union-Find with Path Compression (O(n α(n)) time, O(n) space)
This optimized strategy uses a Union-Find structure combined with subtree traversal. The core observation: only nodes along the path from the node containing genetic value 1 up to the root can have answers greater than 1. All other subtrees are missing value 1. Starting from the node that contains value 1, move upward toward the root while progressively marking nodes in each subtree as visited using DFS. Each visited genetic value is unioned with the next value, effectively tracking the next available missing integer using path compression. The smallest missing value for the current subtree is found by querying the next unconnected number. This avoids rebuilding sets and ensures near-linear complexity.
The solution heavily relies on properties of tree traversal. Subtrees can be explored independently, and the genetic value constraint allows efficient detection of missing integers once values are marked.
Recommended for interviews: The Union-Find optimization is the approach most interviewers expect for this Hard problem. It demonstrates deeper insight into subtree structure and missing-number tracking. Implementing the DFS + set approach first can show your reasoning process, but the optimized Union-Find method proves you can reduce redundant work and reach near-linear complexity.
This approach utilizes depth-first search (DFS) to traverse the tree and a set to track genetic values in each subtree. By doing this, you can efficiently identify the smallest missing genetic value.
The code employs DFS to explore each subtree and gather genetic values using a set. The aim is to determine the smallest missing genetic value in each subtree by iterating through possible genetic numbers and checking their presence.
The time complexity is O(n^2), predominantly due to the iterative process over the genetic values up to n+1 in each subtree DFS traversal. The space complexity is O(n) utilized by the data structures to track children and visited nodes.
This method uses a union-find data structure with path compression to efficiently detect the smallest missing genetic value. It focuses on connectivity and quick access to elements in subtrees.
This Java code integrates a DFS traversal with a union-find approach to efficiently trace and flag seen genetic values using an array. This assists in promptly identifying the smallest missing number.
The time complexity is O(n log n) due to union-find operations with path compression, while space complexity is also O(n) due to auxiliary storage usage.
We notice that each node has a unique gene value, so we only need to find the node idx with gene value 1, and all nodes except for those on the path from node idx to the root node 0 have an answer of 1.
Therefore, we initialize the answer array ans to [1,1,...,1], and our focus is on finding the answer for each node on the path from node idx to the root node 0.
We can start from node idx and use depth-first search to mark the gene values that appear in the subtree rooted at idx, and record them in the array has. During the search process, we use an array vis to mark the visited nodes to prevent repeated visits.
Next, we start from i=2 and keep looking for the first gene value that has not appeared, which is the answer for node idx. Here, i is strictly increasing, because the gene values are unique, so we can always find a gene value that has not appeared in [1,..n+1].
Then, we update the answer for node idx, i.e., ans[idx]=i, and update idx to its parent node to continue the above process until idx=-1, which means we have reached the root node 0.
Finally, we return the answer array ans.
The time complexity is O(n), and the space complexity is O(n). Here, n is the number of nodes.
| Approach | Complexity |
|---|---|
| DFS with a Set to Track Genetic Values | The time complexity is O(n^2), predominantly due to the iterative process over the genetic values up to n+1 in each subtree DFS traversal. The space complexity is O(n) utilized by the data structures to track children and visited nodes. |
| Union-Find with Path Compression | The time complexity is O(n log n) due to union-find operations with path compression, while space complexity is also O(n) due to auxiliary storage usage. |
| DFS | — |
| Approach | Time | Space | When to Use |
|---|---|---|---|
| DFS with Set Tracking | O(n^2) worst case | O(n) | Good for understanding subtree aggregation and quick prototyping |
| Union-Find with Path Compression | O(n α(n)) | O(n) | Optimal solution for large trees and interview settings |
LeetCode 2003. Smallest Missing Genetic Value in Each Subtree • Happy Coding • 1,533 views views
Watch 6 more video solutions →Practice Smallest Missing Genetic Value in Each Subtree with our built-in code editor and test cases.
Practice on FleetCodePractice this problem
Open in Editor