There is a rooted tree consisting of n nodes numbered 0 to n - 1. Each node's number denotes its unique genetic value (i.e. the genetic value of node x is x). The genetic difference between two genetic values is defined as the bitwise-XOR of their values. You are given the integer array parents, where parents[i] is the parent for node i. If node x is the root of the tree, then parents[x] == -1.
You are also given the array queries where queries[i] = [nodei, vali]. For each query i, find the maximum genetic difference between vali and pi, where pi is the genetic value of any node that is on the path between nodei and the root (including nodei and the root). More formally, you want to maximize vali XOR pi.
Return an array ans where ans[i] is the answer to the ith query.
Example 1:
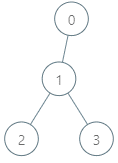
Input: parents = [-1,0,1,1], queries = [[0,2],[3,2],[2,5]] Output: [2,3,7] Explanation: The queries are processed as follows: - [0,2]: The node with the maximum genetic difference is 0, with a difference of 2 XOR 0 = 2. - [3,2]: The node with the maximum genetic difference is 1, with a difference of 2 XOR 1 = 3. - [2,5]: The node with the maximum genetic difference is 2, with a difference of 5 XOR 2 = 7.
Example 2:
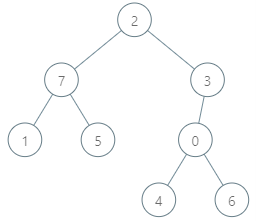
Input: parents = [3,7,-1,2,0,7,0,2], queries = [[4,6],[1,15],[0,5]] Output: [6,14,7] Explanation: The queries are processed as follows: - [4,6]: The node with the maximum genetic difference is 0, with a difference of 6 XOR 0 = 6. - [1,15]: The node with the maximum genetic difference is 1, with a difference of 15 XOR 1 = 14. - [0,5]: The node with the maximum genetic difference is 2, with a difference of 5 XOR 2 = 7.
Constraints:
2 <= parents.length <= 1050 <= parents[i] <= parents.length - 1 for every node i that is not the root.parents[root] == -11 <= queries.length <= 3 * 1040 <= nodei <= parents.length - 10 <= vali <= 2 * 105Problem Overview: You are given a rooted tree where each node represents a genetic value equal to its index. For every query [node, val], compute the maximum val XOR x where x is any ancestor of node (including the node itself). The challenge is answering many XOR queries efficiently while traversing the tree.
Approach 1: DFS with Trie (Time: O((n + q) * logV), Space: O(n * logV))
This approach performs a depth-first search over the tree while maintaining a binary Trie of the values on the current root-to-node path. When DFS enters a node, insert the node's value into the Trie. For every query attached to that node, compute the maximum XOR with val by greedily choosing the opposite bit at each level of the Trie (a classic bit manipulation trick). After processing children, remove the node value when backtracking so the Trie always reflects the active ancestor path.
The key insight: at any moment during DFS, the Trie contains exactly the ancestors of the current node. That means each query becomes a standard "maximum XOR with set" operation. Since values are bounded by node indices, the Trie depth is about 18–20 bits, making each insert, delete, and query very fast.
Approach 2: Preprocessing with Persistent Trie (Time: O((n + q) * logV), Space: O(n * logV))
This approach builds a persistent binary Trie for each node representing the set of values from the root to that node. Instead of modifying a single Trie during DFS, every insertion creates a new version that shares most of its structure with the previous one. The version for a node is derived from its parent's version with the node value inserted.
Each query simply uses the persistent Trie version corresponding to the queried node and performs the same maximum XOR lookup. Because nodes share structure between versions, memory growth stays manageable while avoiding insert/remove operations during traversal.
The persistent structure is especially useful when queries are independent of traversal order or when you want immutable versions of the ancestor sets. The lookup process is identical to a standard maximum XOR search in a binary Trie.
Recommended for interviews: DFS with Trie is the approach most interviewers expect. It demonstrates understanding of tree traversal, dynamic data structures, and XOR optimization. The persistent Trie version shows deeper knowledge of functional data structures and is more common in competitive programming or advanced system design discussions.
This approach uses a depth-first search (DFS) combined with a Trie data structure. The Trie is used to exploit the properties of XOR operations in order to efficiently find the maximum genetic difference for each query.
By traversing from the root to the queries' nodes, we can introduce the nodes into the trie as we traverse the DFS path. At each node where a query is made, we use the trie to efficiently get the maximum XOR with the given value by traversing the trie, looking for bits that will maximize the XOR value.
This C++ solution builds a Trie to track numbers available from the root to the node's path and uses a DFS to traverse the tree. As each node is visited, it is added to the Trie. At each query node, it calculates the maximum XOR by traversing the Trie. Before backtracking in DFS, it erases the node from the Trie. This ensures the current node's number is only available to the queries along its path.
C++
Java
Python
JavaScript
C#
Time Complexity: O(N * 32 + Q * 32) ~ O(N + Q), where N is the number of nodes and Q is the number of queries. The factor 32 corresponds to the max bit length for inserting, deleting, and querying in the Trie.
Space Complexity: O(32 * N), related to the Trie size built during DFS traversal.
This approach involves using a persistent Trie to support efficient query processing by maintaining versions of the Trie for each node in the tree. It leverages the persistence to avoid repeated computations in individual DFS traversals.
By maintaining versions of the Trie, we can avoid the need to insert and remove nodes dynamically within a single traversal, aiming to optimize the retrieval process of the maximum XOR value for varied inputs.
By using a persistent Trie data structure, each node in the tree can create its own version of the Trie by adding itself to the most recent Trie version. During DFS, nodes do not need to be added or removed but instead create a new version as they are visited. Each query is processed using the specific version of the Trie that corresponds to the path in the underlying tree structure from the root to the node being queried. This approach allows the data structure to handle queries more efficiently by using optimized retrieval from pre-built trie versions, enhancing performance compared to dynamic insertion and erasure.
C++
Java
Python
JavaScript
C#
Time Complexity: O(N * 32 + Q * 32) ~ O(N + Q), considering node and query bit operations.
Space Complexity: The use of a persistent Trie could slightly increase space usage, maintaining different Trie states.
| Approach | Complexity |
|---|---|
| Approach 1: DFS with Trie | Time Complexity: O(N * 32 + Q * 32) ~ O(N + Q), where N is the number of nodes and Q is the number of queries. The factor 32 corresponds to the max bit length for inserting, deleting, and querying in the Trie. Space Complexity: O(32 * N), related to the Trie size built during DFS traversal. |
| Approach 2: Preprocessing with Persistent Trie | Time Complexity: O(N * 32 + Q * 32) ~ O(N + Q), considering node and query bit operations. Space Complexity: The use of a persistent Trie could slightly increase space usage, maintaining different Trie states. |
| Approach | Time | Space | When to Use |
|---|---|---|---|
| DFS with Trie | O((n + q) * logV) | O(n * logV) | Best general solution. Efficient when queries are processed during DFS traversal. |
| Preprocessing with Persistent Trie | O((n + q) * logV) | O(n * logV) | Useful when immutable versions of ancestor sets are needed or queries are processed independently. |
LeetCode 1938. Maximum Genetic Difference Query • Happy Coding • 1,383 views views
Watch 4 more video solutions →Practice Maximum Genetic Difference Query with our built-in code editor and test cases.
Practice on FleetCode