There is a family tree rooted at 0 consisting of n nodes numbered 0 to n - 1. You are given a 0-indexed integer array parents, where parents[i] is the parent for node i. Since node 0 is the root, parents[0] == -1.
There are 105 genetic values, each represented by an integer in the inclusive range [1, 105]. You are given a 0-indexed integer array nums, where nums[i] is a distinct genetic value for node i.
Return an array ans of length n where ans[i] is the smallest genetic value that is missing from the subtree rooted at node i.
The subtree rooted at a node x contains node x and all of its descendant nodes.
Example 1:
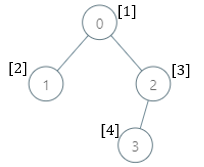
Input: parents = [-1,0,0,2], nums = [1,2,3,4] Output: [5,1,1,1] Explanation: The answer for each subtree is calculated as follows: - 0: The subtree contains nodes [0,1,2,3] with values [1,2,3,4]. 5 is the smallest missing value. - 1: The subtree contains only node 1 with value 2. 1 is the smallest missing value. - 2: The subtree contains nodes [2,3] with values [3,4]. 1 is the smallest missing value. - 3: The subtree contains only node 3 with value 4. 1 is the smallest missing value.
Example 2:
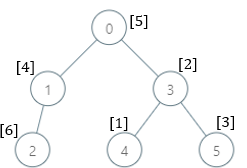
Input: parents = [-1,0,1,0,3,3], nums = [5,4,6,2,1,3] Output: [7,1,1,4,2,1] Explanation: The answer for each subtree is calculated as follows: - 0: The subtree contains nodes [0,1,2,3,4,5] with values [5,4,6,2,1,3]. 7 is the smallest missing value. - 1: The subtree contains nodes [1,2] with values [4,6]. 1 is the smallest missing value. - 2: The subtree contains only node 2 with value 6. 1 is the smallest missing value. - 3: The subtree contains nodes [3,4,5] with values [2,1,3]. 4 is the smallest missing value. - 4: The subtree contains only node 4 with value 1. 2 is the smallest missing value. - 5: The subtree contains only node 5 with value 3. 1 is the smallest missing value.
Example 3:
Input: parents = [-1,2,3,0,2,4,1], nums = [2,3,4,5,6,7,8] Output: [1,1,1,1,1,1,1] Explanation: The value 1 is missing from all the subtrees.
Constraints:
n == parents.length == nums.length2 <= n <= 1050 <= parents[i] <= n - 1 for i != 0parents[0] == -1parents represents a valid tree.1 <= nums[i] <= 105nums[i] is distinct.The key observation in #2003 Smallest Missing Genetic Value in Each Subtree is that most subtrees do not contain the value 1. For such nodes, the smallest missing genetic value is automatically 1. The main work therefore focuses on the path from the node containing 1 up to the root.
A practical strategy uses Depth-First Search (DFS) combined with a structure to track visited genetic values. Starting from the node containing 1, gradually expand to its ancestors while marking all values in the explored subtrees. For each step upward, update the smallest missing positive value that has not been seen yet. Efficient tracking can be implemented using boolean arrays or hash sets.
This approach avoids recomputing values for every subtree and processes each node and genetic value at most once. The algorithm runs in roughly O(n) time with O(n) additional space for tracking visited nodes and values.
| Approach | Time Complexity | Space Complexity |
|---|---|---|
| DFS with value tracking from node containing 1 | O(n) | O(n) |
NeetCode
Use these hints if you're stuck. Try solving on your own first.
If the subtree doesn't contain 1, then the missing value will always be 1.
What data structure allows us to dynamically update the values that are currently not present?
Watch expert explanations and walkthroughs
Practice problems asked by these companies to ace your technical interviews.
Explore More ProblemsJot down your thoughts, approach, and key learnings
If a subtree does not contain the genetic value 1, the smallest missing positive value must be 1. This observation significantly reduces the work required because only the subtree containing 1 and its ancestors need detailed processing.
Yes, problems involving trees, DFS, and clever observations about constraints are common in FAANG-style interviews. This question tests understanding of tree traversal, value tracking, and optimization strategies for avoiding repeated work.
A boolean array or hash set is commonly used to track which genetic values have already appeared in explored subtrees. Combined with an adjacency list representation of the tree and DFS traversal, it allows efficient updates of the smallest missing value.
The optimal approach focuses on the node containing value 1 and processes its path to the root. Using DFS and a structure to track seen genetic values, we update the smallest missing value as we expand through relevant subtrees. This prevents redundant calculations and achieves near linear time.